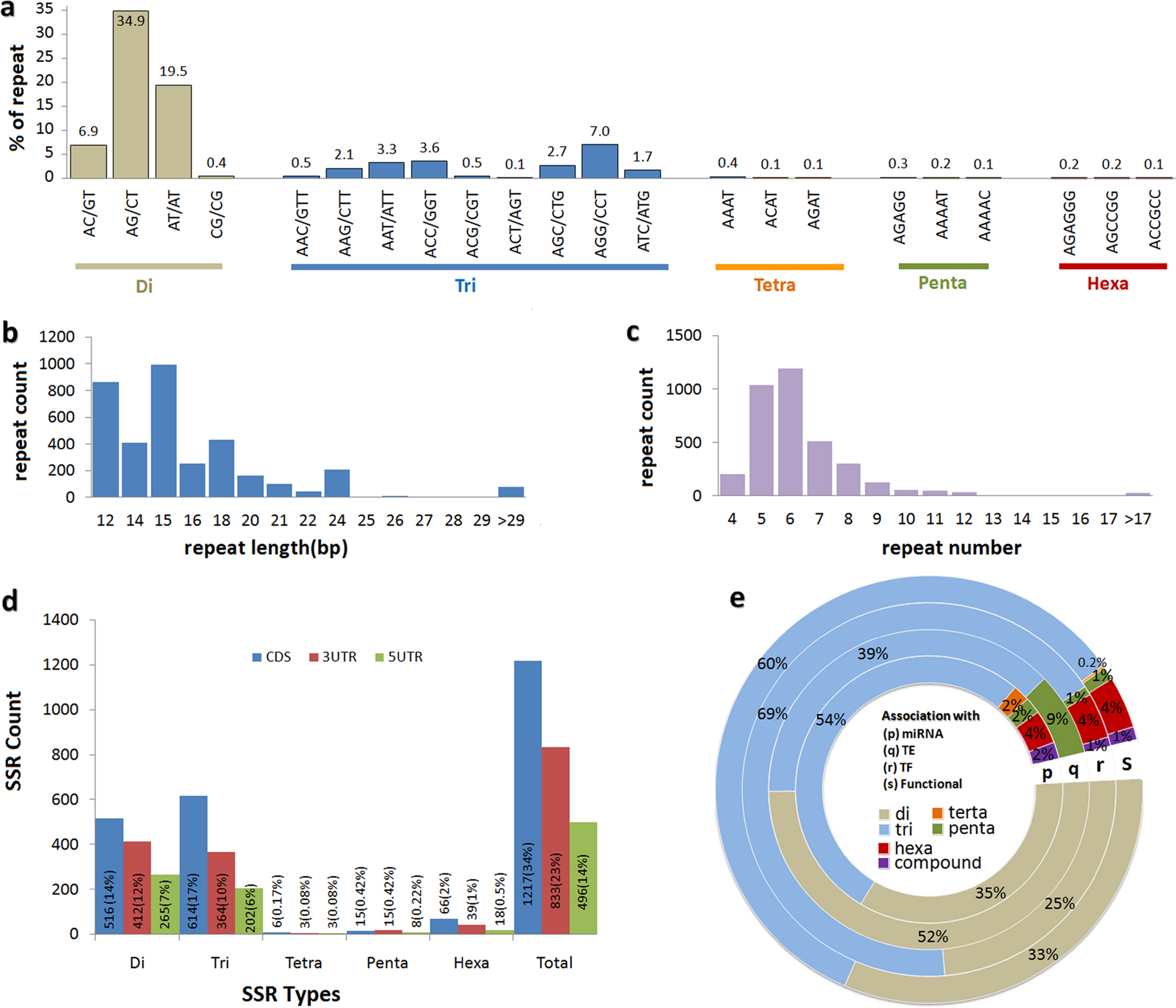
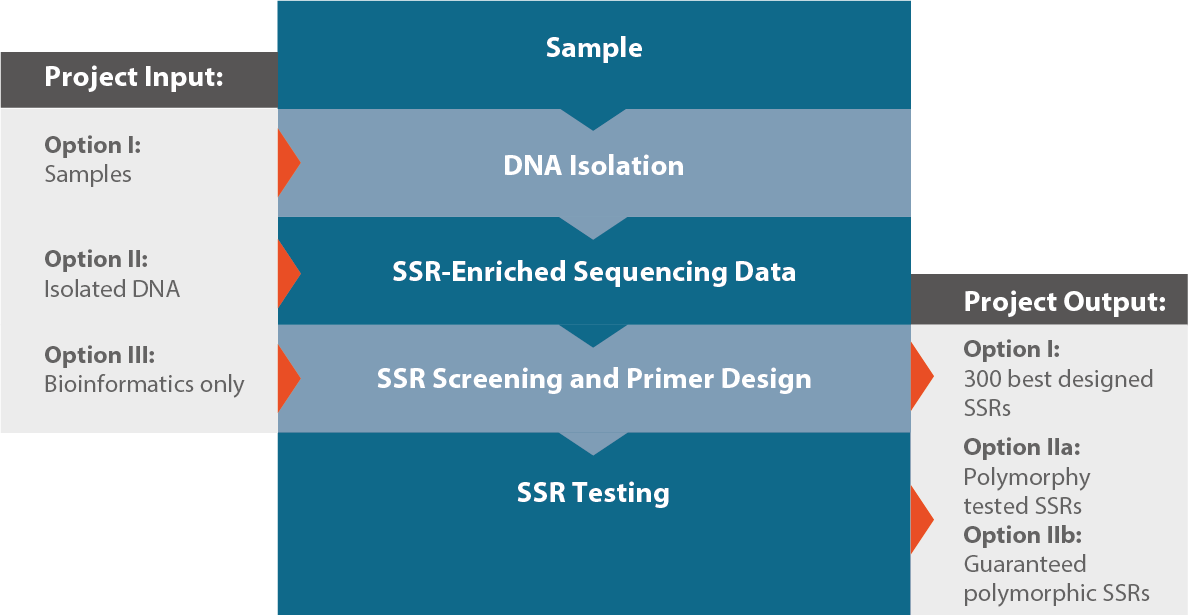
Genes | Free Full-Text | Transferability and Polymorphism of SSR Markers Located in Flavonoid Pathway Genes in Fragaria and Rubus Species
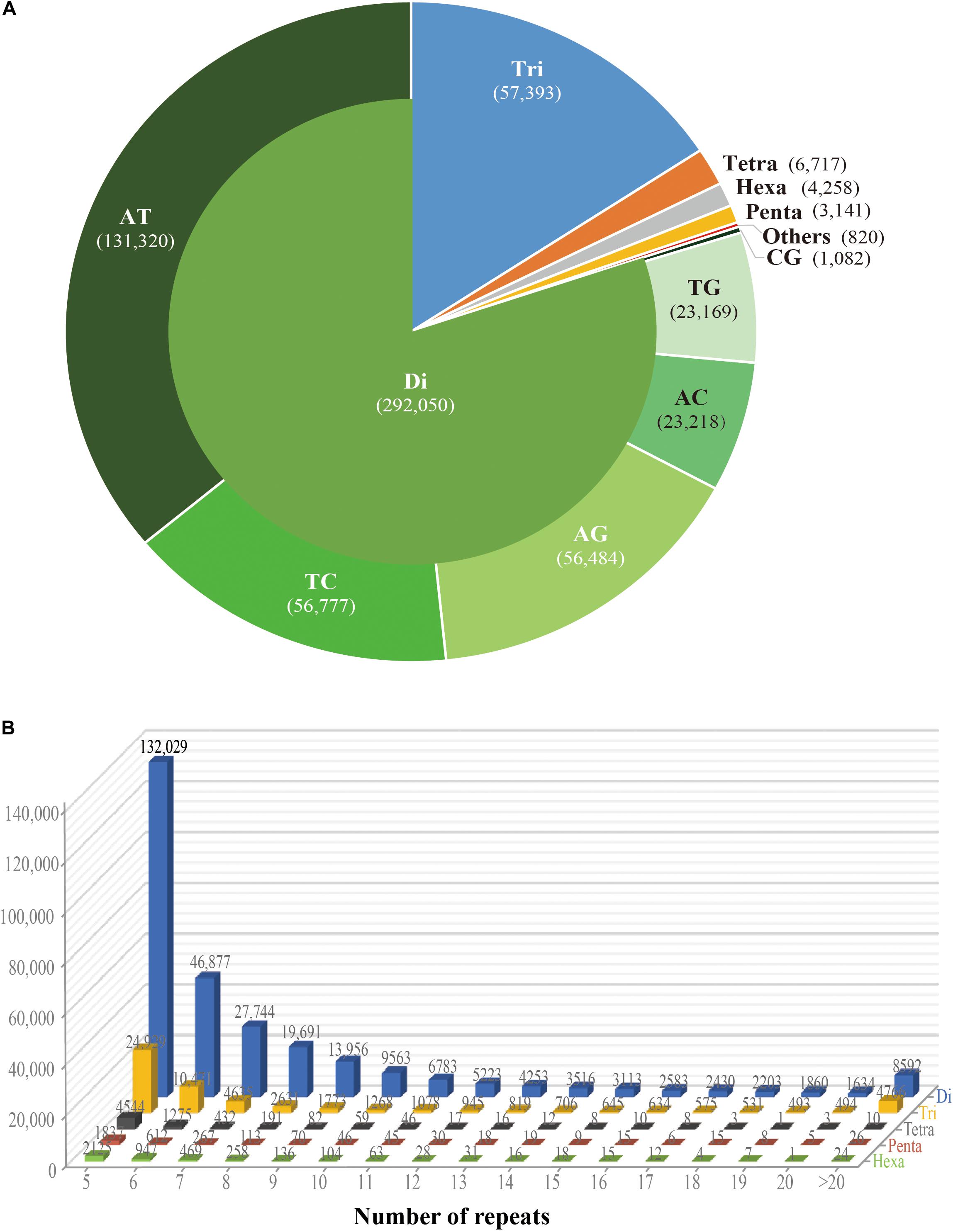
Frontiers | Development of a Large Gene-Associated SSR Marker Set and in-Depth Genetic Characterization in Scarlet Sage
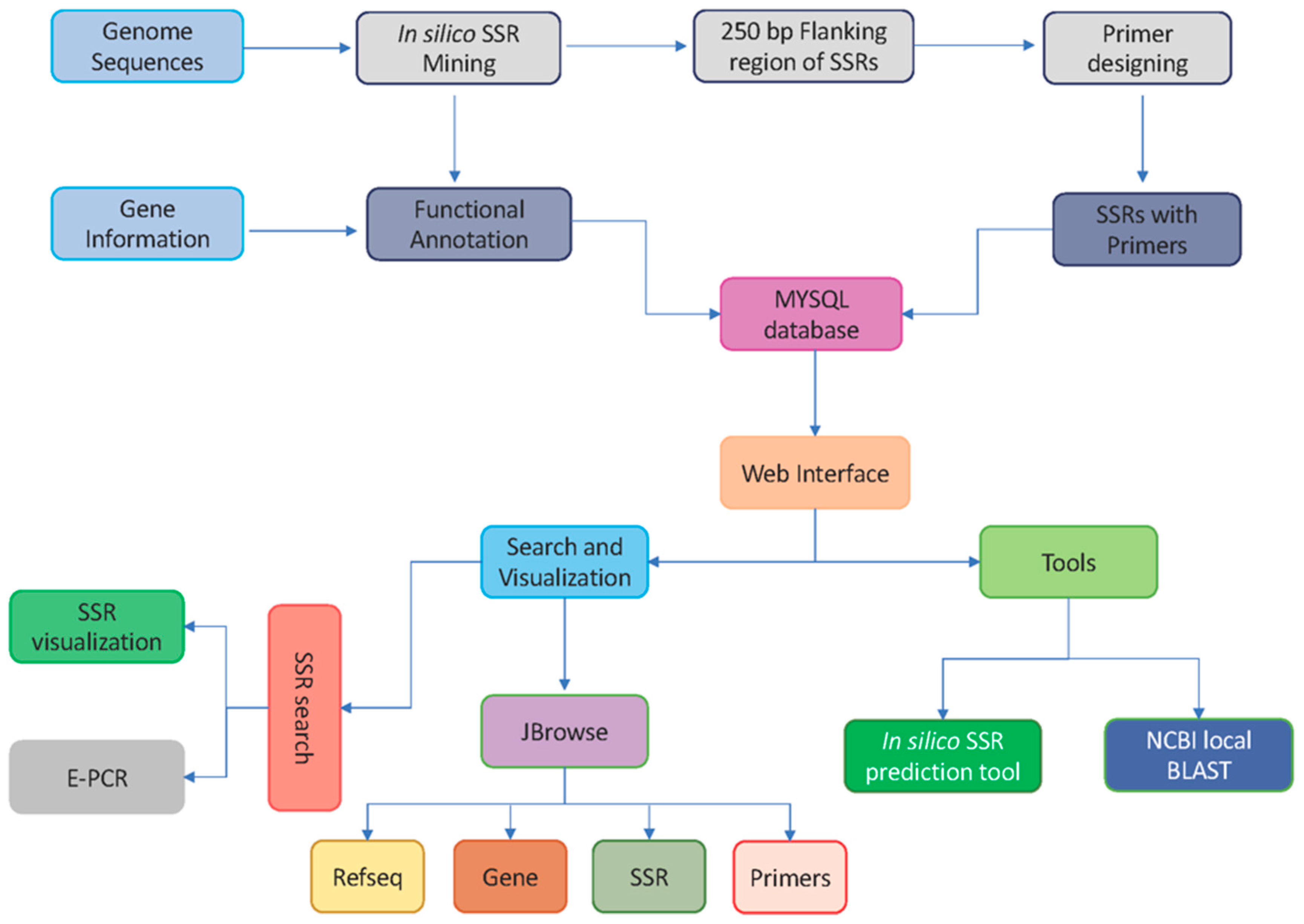
Genes | Free Full-Text | citSATdb: Genome-Wide Simple Sequence Repeat (SSR) Marker Database of Citrus Species for Germplasm Characterization and Crop Improvement

Molecules | Free Full-Text | Mining and Development of Novel SSR Markers Using Next Generation Sequencing (NGS) Data in Plants

Forests | Free Full-Text | Genetic Diversity and Association Analysis among Germplasms of Diospyros kaki in Zhejiang Province Based on SSR Markers
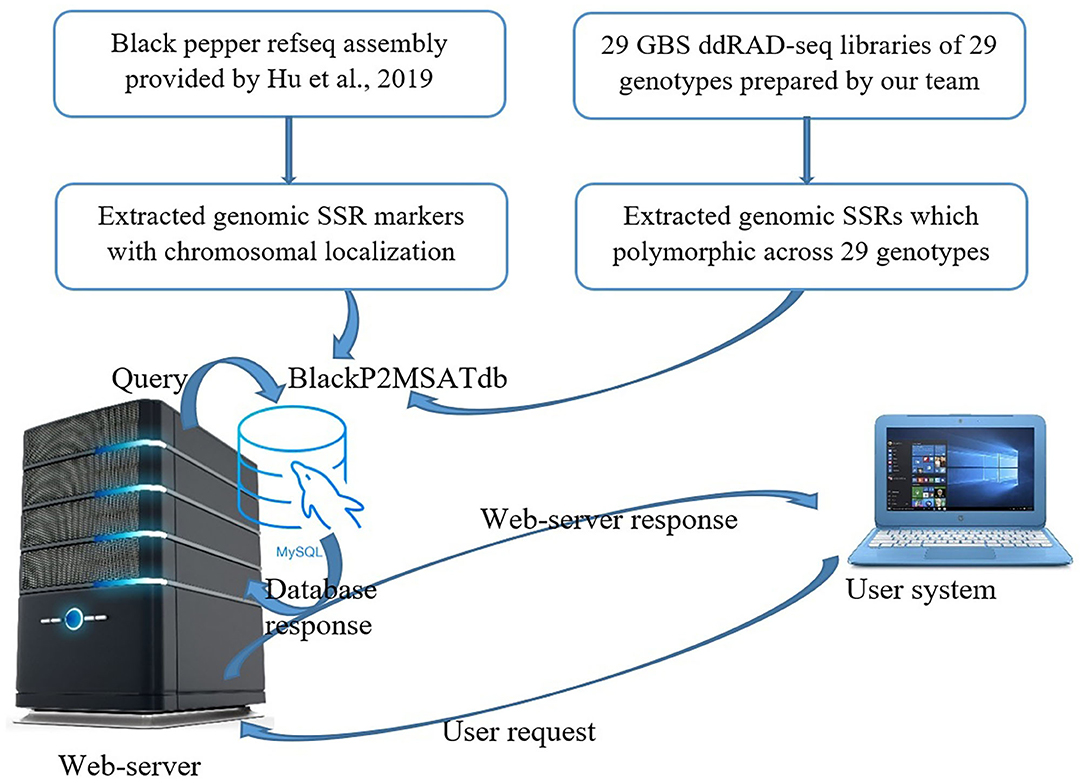
Frontiers | Rapid Genome-Wide Location-Specific Polymorphic SSR Marker Discovery in Black Pepper by GBS Approach
Genome-Wide Characterization of Simple Sequence Repeat (SSR) Loci in Chinese Jujube and Jujube SSR Primer Transferability | PLOS ONE

Genome-wide identification of simple sequence repeats and development of polymorphic SSR markers in swamp eel (Monopterus albus) - Hai-feng Tian, Qiao-mu Hu, Zhong Li, 2021
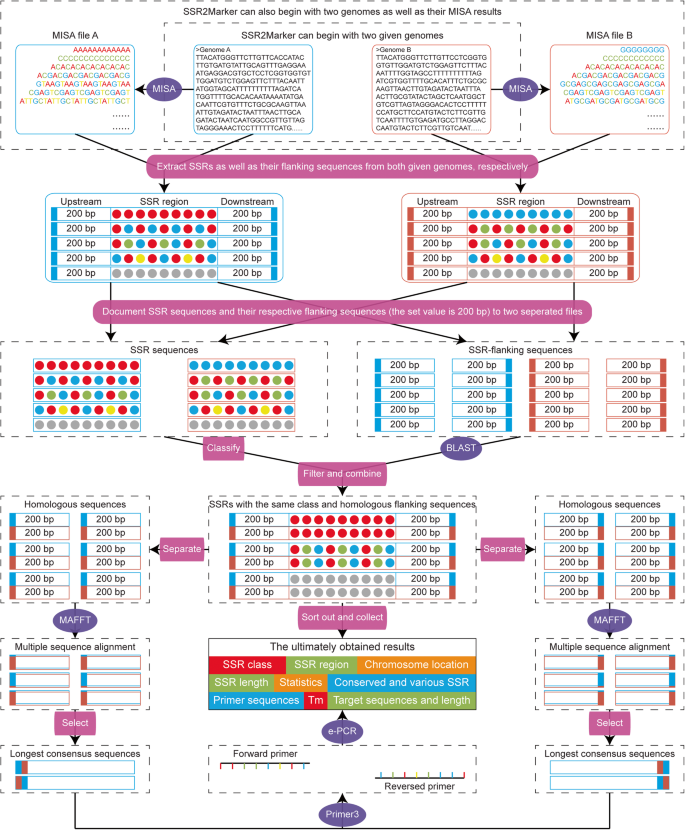
SSR2Marker: an integrated pipeline for identification of SSR markers within any two given genome-scale sequences | Molecular Horticulture | Full Text
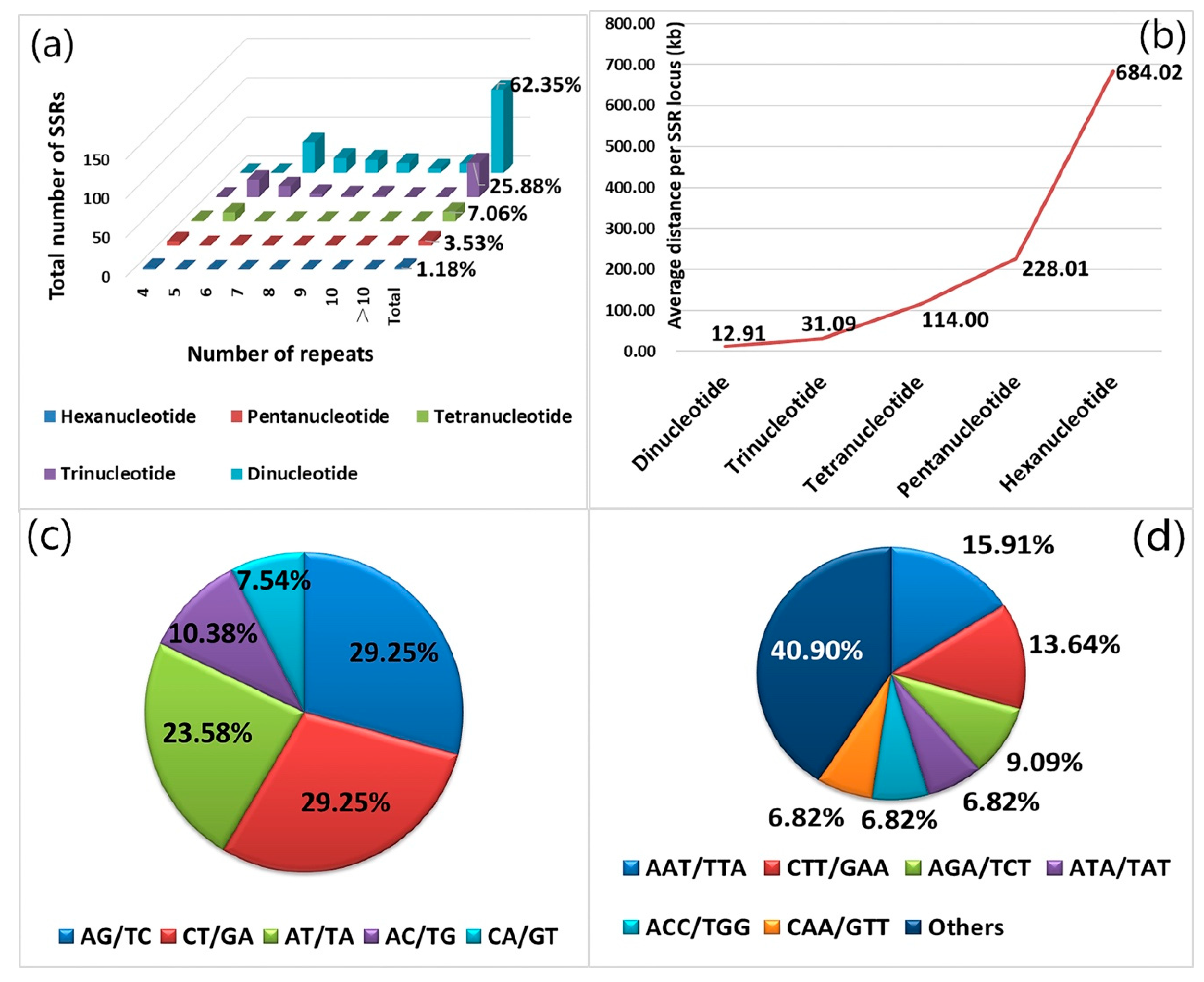
Forests | Free Full-Text | Development and Application of EST-SSR Markers for DNA Fingerprinting and Genetic Diversity Analysis of the Main Cultivars of Black Locust (Robinia pseudoacacia L.) in China
Development and Characterization of Simple Sequence Repeat (SSR) Markers Based on RNA-Sequencing of Medicago sativa and In silico Mapping onto the M. truncatula Genome | PLOS ONE
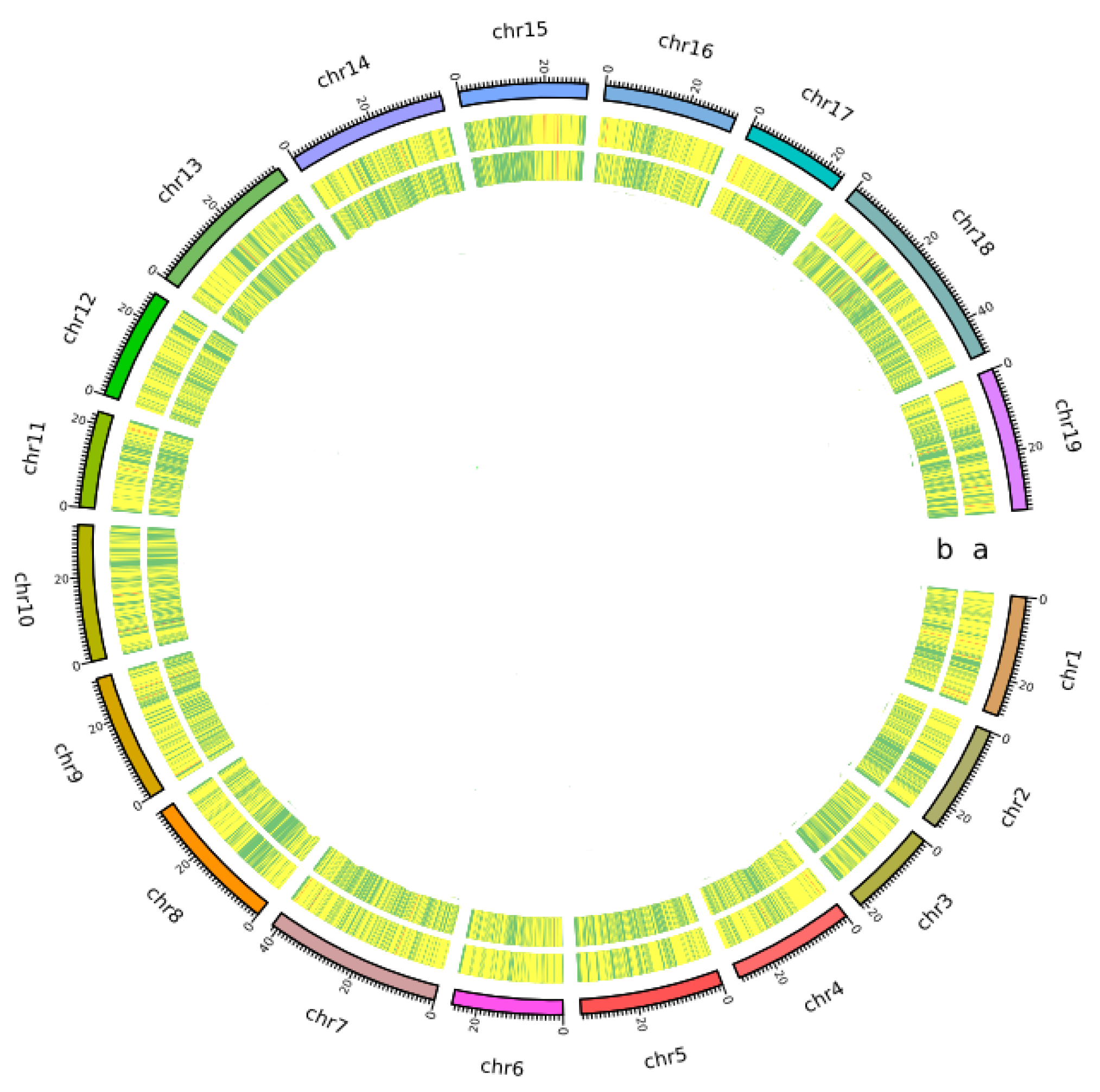
Genes | Free Full-Text | Characterization of Simple Sequence Repeat (SSR) Markers Mined in Whole Grape Genomes
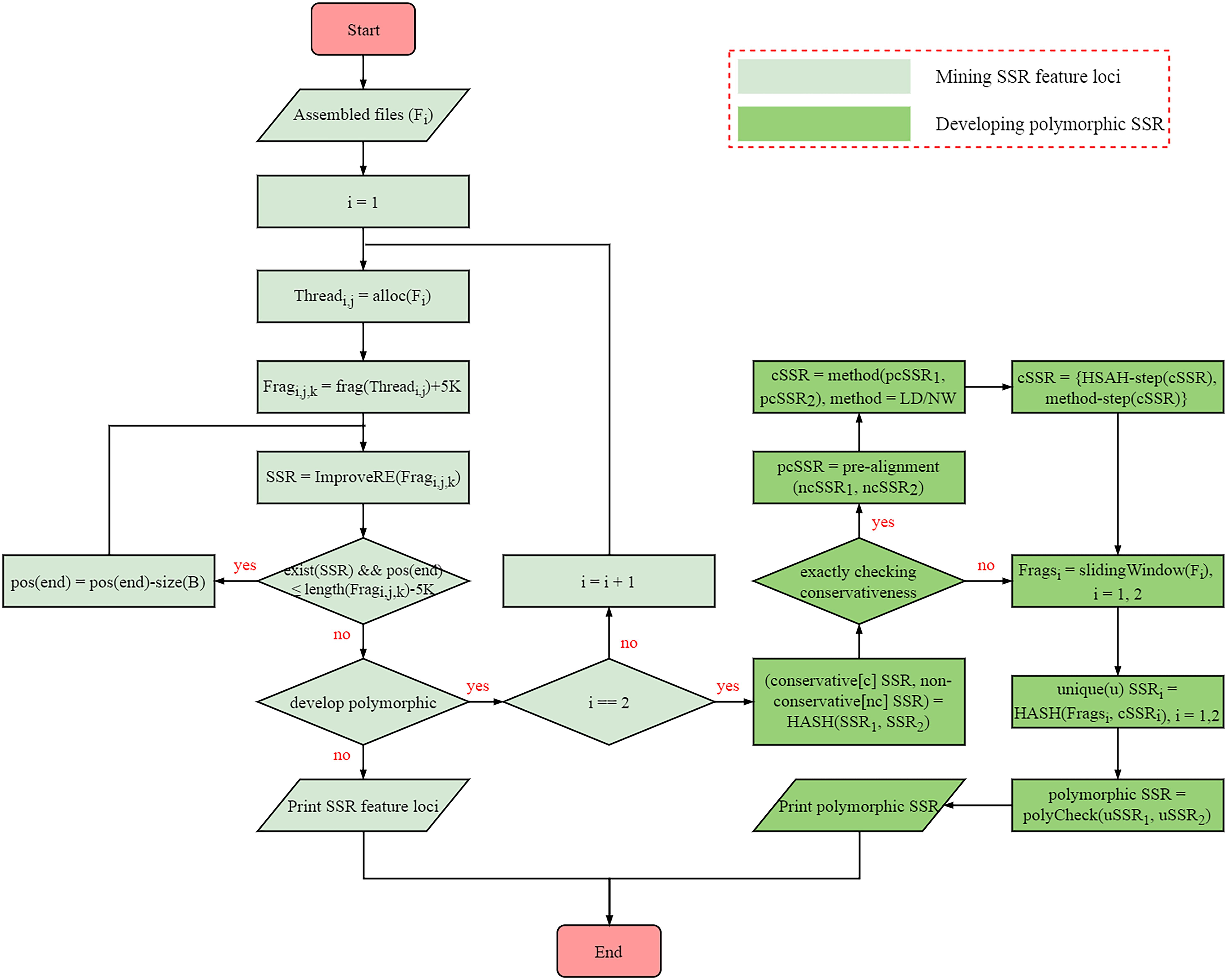
Frontiers | SSRMMD: A Rapid and Accurate Algorithm for Mining SSR Feature Loci and Candidate Polymorphic SSRs Based on Assembled Sequences